About
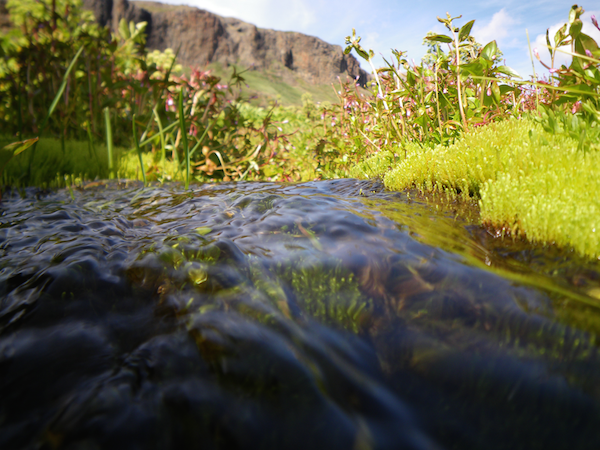
-The Dunn Lab is in the Department of Ecology and Evolutionary Biology at Yale University.
- -We study how evolution has produced a diversity of life. We are interested in learning about the actual history of life on Earth as well as the general properties of evolution that have contributed to these historical patterns. The type of questions we ask require field (marine), laboratory, and computational work. Our research falls into several domains.
CreatureCast - How Siphonophores Grow. A CreatureCast animation by Riley Thompson that explains some of what we have learned about how siphonophores grow. See our medium post for more.
-
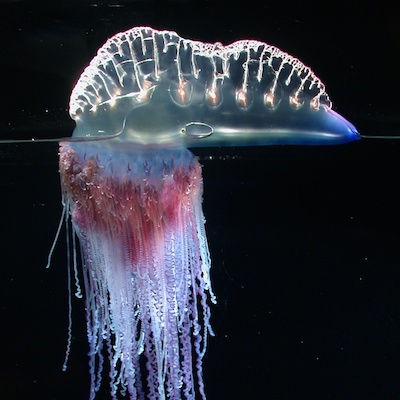

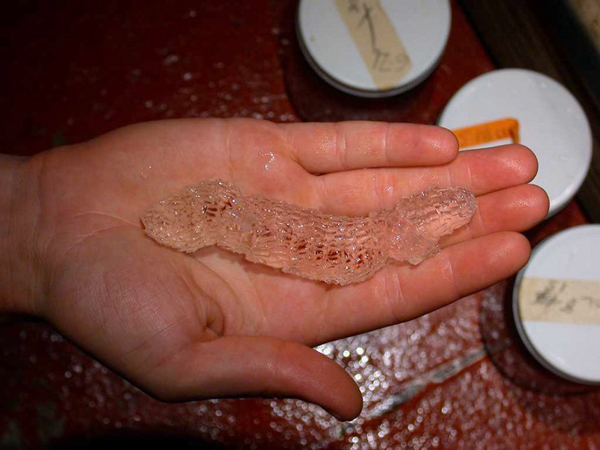
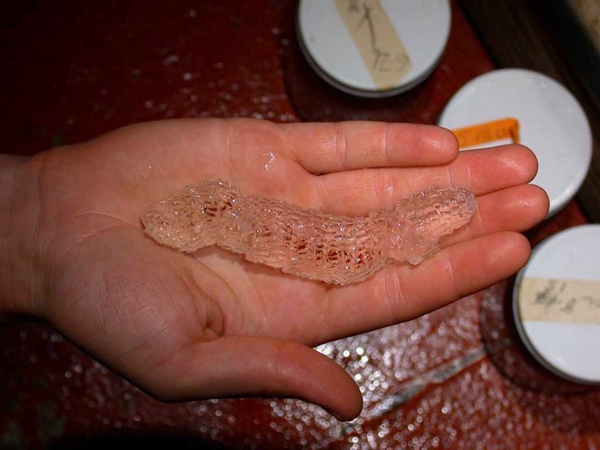
Siphonophores
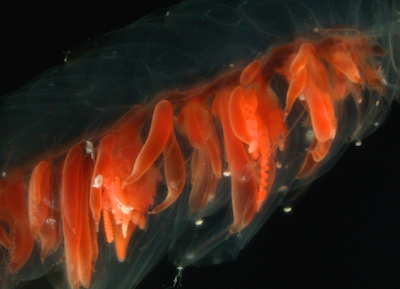
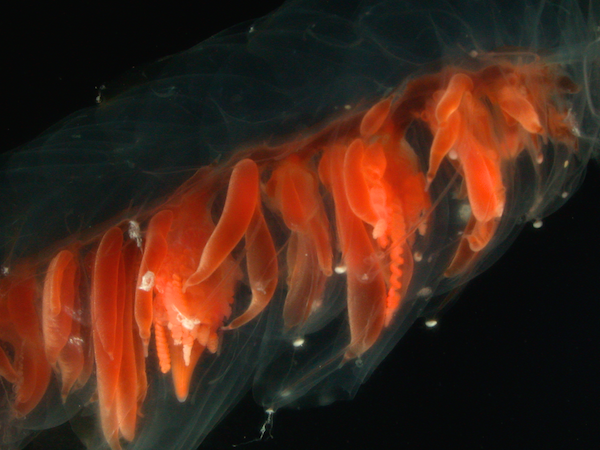
Siphonophores are unique marine animals found throughout the oceans of the world. They are colonial - each siphonophore starts as a single embryo that asexually produces many genetically identical, physiologically integrated bodies. The many bodies in a siphonophore are each specialized for particular functions, such as feeding or swimming. We study the morphology, evolution, and development of siphonophores to understand this unique functional specialization.
-
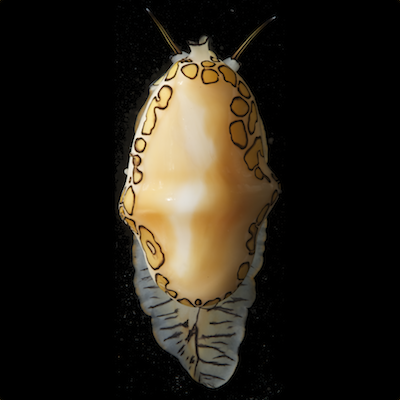

Animal phylogenetics
Phylogenetics is the study of evolutionary relationships. We develop new methods and tools for understanding phylogenetic relationships, and apply these tools to animals to understand the evolution of complex traits. We are particularly interested in deep animal phylogeny, but also work on subgroups of animals including cnidarians and molluscs.
-
Methods and tools
Many of the questions we are work on require new computational methods and tools. We develop and implement new methods, connect existing analysis tools in new ways, and are particularly interested in engineering analyses so that they are open, transparent, reproducible, and easy to extend. Our data analysis tools are available at bitbucket.
-
Evolution of gene expression
We sequence transcriptomes to build phylogenies and to identify genes that are differentially expressed between tissues and developmental stages. We have integrated these two approaches to now study the evolution of differential gene expression. This work provides new ways to understand the relationships between genome and phenotype evolution.
-
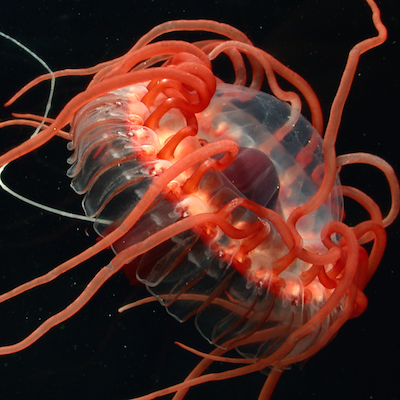

Zoology
In addition to the specific domains highlighted above, we are very interested in the basic biology of animals. Since many of the animals we work on are so poorly known, these studies include descriptions of lifecycles, natural history, development, alpha taxonomy, functional biology, and morphology.